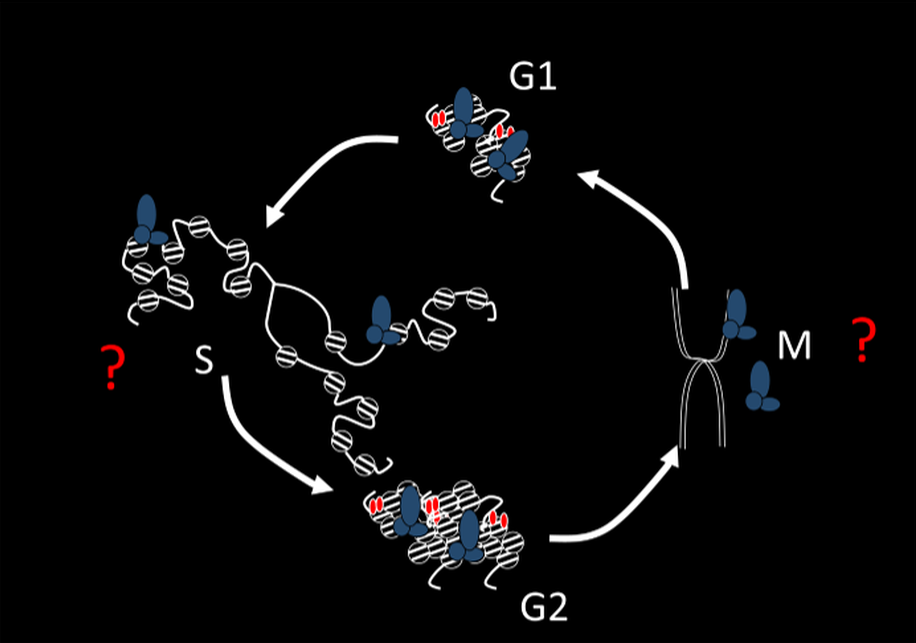
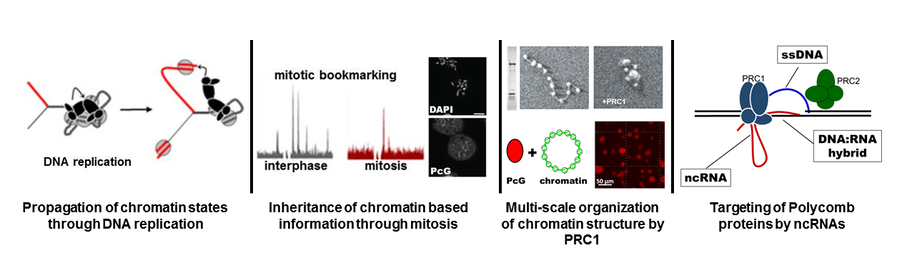
The ability of cells to acquire and maintain stable differentiated states is central to development. Our lab is interested in how the gene expression patterns that define cellular states can be inherited across cell generations and how this might involve heritable changes to chromatin. The idea that chromatin structure carries heritable (epigenetic) information is attractive because of its conceptual simplicity-epigenetic and genetic information are inherited together. However, this idea raises questions about how chromatin features can be inherited through cell cycles since both DNA replication and mitosis disrupt chromatin structure. The goal of our work is to discover the biochemical mechanisms that enable chromatin states to be propagated through DNA replication and mitosis. We address these questions using the Polycomb Group (PcG) proteins of Drosophila melanogaster, and a combination of in vitro reconstitution and in vivo analysis.
HOW ARE CHROMATIN STATES PROPAGATED THROUGH DNA REPLICATION?
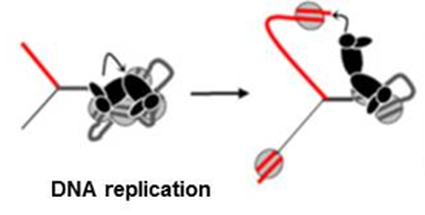
During DNA replication, chromatin–based regulatory information faces two challenges, the disruption caused by passage of the replication machinery, and the two-fold increase in DNA. We aim to understand how of these challenges are met in the PcG system by reconstituting propagation of chromatin states through DNA replication. Current projects involve analyzing the behavior of both PcG proteins and histones during DNA replication, and how chromatin structure is restored after DNA replication.
HOW ARE CHROMATIN STATES PROPAGATED THROUGH MITOSIS?
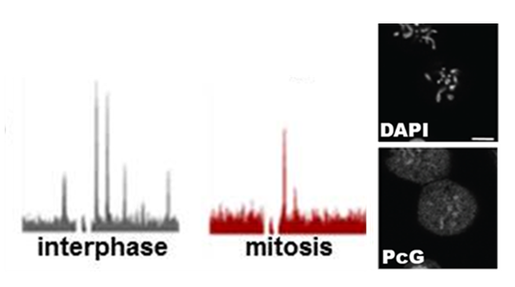
For transcription states to be preserved through cell division, chromatin structures must also survive mitosis, when chromosome structure is altered and many factors are released from chromosomes. We recently identified two classes of PcG protein binding sites in Drosophila cells, one of which retains PcG proteins in mitosis. The features of the sites that persist in mitosis suggest they could have a role in higher order organization of chromatin. Current work aims to determine how and why PcG proteins are lost from some sites in mitosis but not others, and how PcG proteins are recruited back to their target genes as cells exit mitosis. We are carrying out an RNAi screen to identify factors that control Polycomb protein binding to chromosomes in mitosis, and using AP-MS to identify mitosis specific Polycomb protein interactors.
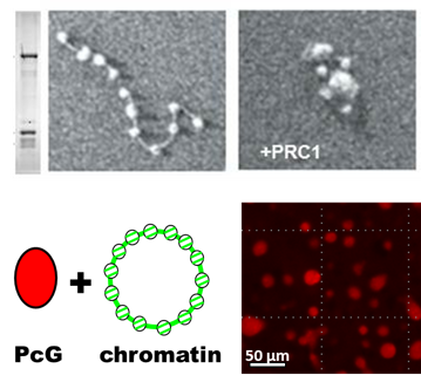
HOW DO POLYCOMB PROTEINS ORGANIZE CHROMATIN?
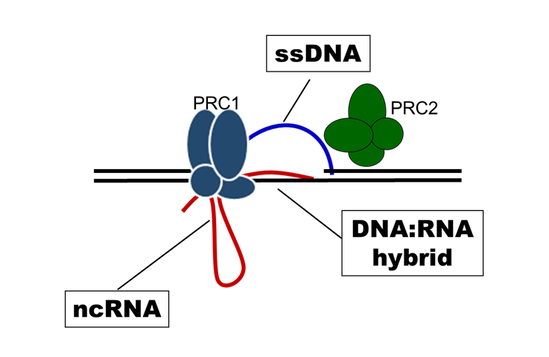
Polycomb proteins organize chromatin into unique large-scale domains in vivo. We aim to reconstitute Polycomb protein-mediated chromatin organization in vitro to understand how this occurs. We are investigating the role of the SAM polymerization domain of Polyhomeotic (with Dr. Chongwoo Kim), and phase separation in chromatin organization. We are also using Electron Microscopy and cross-linking mass spectrometry to understand how PRC1 organizes chromatin more locally.